Nuclear pores in their natural context
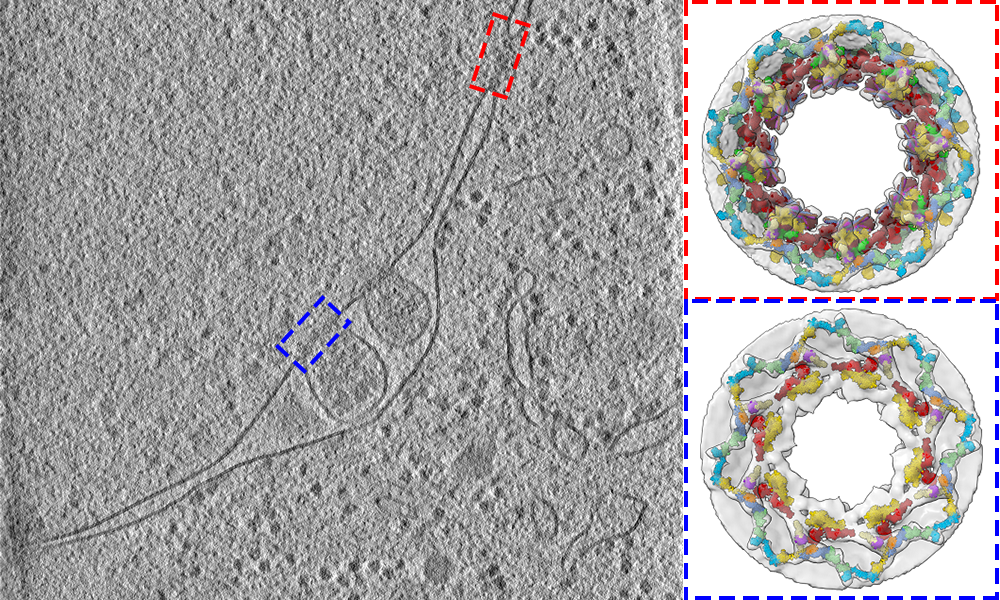
MPIB scientists have contributed to deciphering the 3D structure of the nuclear pore of baker's yeast cells.
Research group leader Boris Pfander and his team from the Max Planck Institute of Biochemistry, together with colleagues from the Max Planck Institute of Biophysics in Frankfurt am Main and the EMBL in Heidelberg, have investigated the 3D structure of nuclear pores in budding yeast (Saccharomyces cerevisiae). Their results, published in Nature, reveal the architecture of the nuclear pore complex in intact cells and broaden our understanding of crucial processes in life.
Nuclear pores are a highly complex assembly of proteins. Hundreds of them are embedded in the double membrane that surrounds and protects the cell’s nucleus. They act as a gateway that regulates the entry and exit of molecules. An important function of nuclear pores is to regulate the export of a molecule called messenger RNA (mRNA) from the nucleus into the surrounding cell – the cytoplasm – where it delivers instructions for the assembly of proteins. Revealing the architecture Now the scientists appreciate better how the nuclear pore works in its native context, how it is maintained and recycled. The study provides a detailed structural description of the three protein rings that make up the nuclear pore, known as the cytoplasmic, nuclear, and inner rings. To show how these rings are arranged in cells, the researchers used a combination of cell biology, computational modelling, and in-cell cryo-electron tomography: an imaging technique, that is used to produce high-resolution 3D views of the molecular landscape inside a cell. This led to fundamental new insights. The scientist found out that the 3D configuration of the cytoplasmic ring accommodates the path of mRNA export.
